-Search query
-Search result
Showing 1 - 50 of 95 items for (author: bernat & a)

EMDB-18520: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18521: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18522: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-18523: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo2: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM
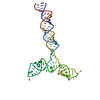
PDB-8qo3: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo4: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

PDB-8qo5: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
Method: single particle / : Moura TR, Purta E, Bernat A, Baulin E, Mukherjee S, Bujnicki JM

EMDB-27177: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

PDB-8d48: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein
Method: single particle / : Abernathy ME, Barnes CO

EMDB-14888: 
CM01B in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14889: 
CM05A1 in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14890: 
CM02A in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14891: 
CM04F in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14892: 
CM03B in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14893: 
CM05E in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14894: 
CM05L in complex with conM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14895: 
NHP MCM02 wk26 N355/N289 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14896: 
NHP MCM04 wk26 base pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14897: 
NHP MCM04 wk26 N611 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14898: 
NHP MCM05 wk26 V1V2V3 and base pAbs in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14899: 
NHP MCM06 wk26 base and N611 pAbs in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14900: 
NHP MCM02 wk26 N355/N289 pAb in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14901: 
NHP CCK046 wk70 base pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14902: 
NHP MCM06 wk70 N355/N289 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14903: 
NHP MCM06 wk70 N611 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14904: 
NHP MCM04 wk70 V1V2V3 pAb in complex with ConM SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14905: 
NHP MCM03 wk70 base pAb in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14906: 
NHP MCM04 wk70 N611 pAb in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14907: 
NHP MCM04 wk70 V1V2V3 pAb in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB

EMDB-14908: 
NHP MCM06 wk70 N355/N289 pAb in complex with ConS SOSIP
Method: single particle / : van Schooten J, Ward AB
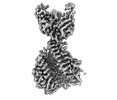
EMDB-27939: 
Structure of the human ACE2 receptor in complex with antibody Fab fragment, 05B04
Method: single particle / : Barnes CO
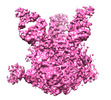
EMDB-24853: 
Modified BG505 SOSIP-based immunogen RC1 in complex with elicited mAb Ab283MUR Fab and 8ANC195 Fab
Method: single particle / : Abernathy ME, Bjorkman PJ

EMDB-24854: 
Modified BG505 SOSIP-based immunogen RC1 in complex with elicited mAb Ab1170NHP Fab and 8ANC195 Fab
Method: single particle / : Abernathy ME, Bjorkman PJ

EMDB-24855: 
Modified BG505 SOSIP-based immunogen RC1 in complex with Rabbit 2249 post-Prime polyclonal Fabs
Method: single particle / : Barnes CO, DeLaitsch AT, Bjorkman PJ

EMDB-24856: 
Modified BG505 SOSIP-based immunogen RC1 in complex with Rabbit 2249 post-Boost 1 polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24857: 
Modified BG505 SOSIP-based immunogen RC1 in complex with Rabbit 2249 post-Boost 2 polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24858: 
Modified BG505 SOSIP-based immunogen RC1 in complex with Rabbit 2249 post-Boost 3 polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24859: 
Modified BG505 SOSIP-based immunogen RC1 in complex with Rabbit 2249 post-Boost 4 polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24860: 
Modified BG505 SOSIP-based immunogen RC1 in complex with NHP 1 post-Prime polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24861: 
Modified BG505 SOSIP-based immunogen RC1 in complex with NHP 1 post-Boost 1 polyclonal Fabs
Method: single particle / : DeLaitsch AT, Barnes CO, Bjorkman PJ

EMDB-24072: 
Ab1245 Fab in complex with BG505 SOSIP.664 and 8ANC195 Fab
Method: single particle / : Abernathy ME, Bjorkman PJ

PDB-7mxe: 
Ab1245 Fab in complex with BG505 SOSIP.664 and 8ANC195 Fab
Method: single particle / : Abernathy ME, Bjorkman PJ

EMDB-23393: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C601
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23394: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C603
Method: single particle / : Yang Z, Barnes CO, Bjorkman PJ

EMDB-23395: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C643
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23396: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C663
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23397: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C666
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23398: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C669
Method: single particle / : Barnes CO, Bjorkman PJ

EMDB-23399: 
Structure of the SARS-CoV-2 spike trimer in complex with mRNA vaccine induced neutralizing antibody C670
Method: single particle / : Yang Z, Barnes CO, Bjorkman PJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
